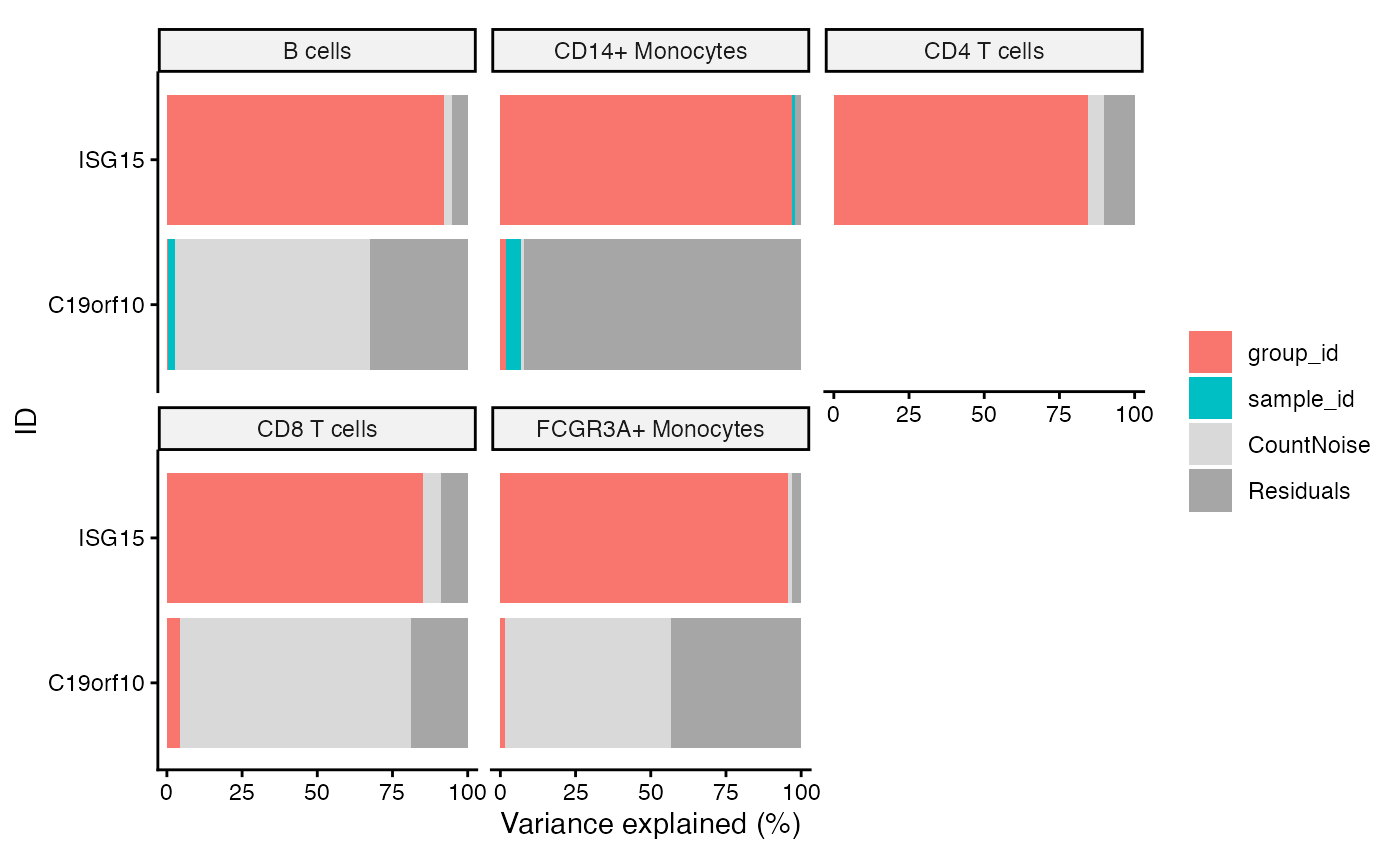
Bar plot of variance fraction for each gene and each variable
Usage
plotPercentBars(
x,
col = c(ggColorHue(base::ncol(x) - 4), "grey85", "grey65"),
ncol = 3,
cluster_ids = unique(x[["cluster_id"]]),
...
)
# S4 method for class 'data.frame'
plotPercentBars(
x,
col = c(ggColorHue(base::ncol(x) - 4), "grey85", "grey65"),
ncol = 3,
cluster_ids = unique(x[["cluster_id"]]),
...
)Arguments
- x
object returned by
fitVarPart()- col
vector of colors
- ncol
number of columns in the plot
- cluster_ids
which cell types to plot
- ...
additional arguments
Examples
library(SingleCellExperiment)
# Load example data
data(example_sce, package="muscat")
sce <- example_sce
# Compute library size for each cell
sce$libSize <- colSums(counts(sce))
# Specify regression formula and cell annotation
form <- ~ group_id + (1|sample_id)
fit <- lucida(sce, form, "cluster_id", verbose=FALSE)
#> B cells
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:01
#> CD14+ Monocytes
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:01
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:01
#> CD4 T cells
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:01
#> CD8 T cells
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:01
#> FCGR3A+ Monocytes
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:01
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:01
# Model with only intercept and random effect
form <- ~ (1|sample_id)
fit.null <- lucida(sce, form, "cluster_id", verbose=FALSE)
#> B cells
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#> CD14+ Monocytes
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#> CD4 T cells
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#> CD8 T cells
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#> FCGR3A+ Monocytes
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
# Variance partitioning analysis
vp <- fitVarPart(fit, fit.null)
# Bar plots of a subset of genes
library(dplyr)
vp %>%
sortCols %>%
filter(ID %in% c('ISG15', 'C19orf10')) %>%
plotPercentBars