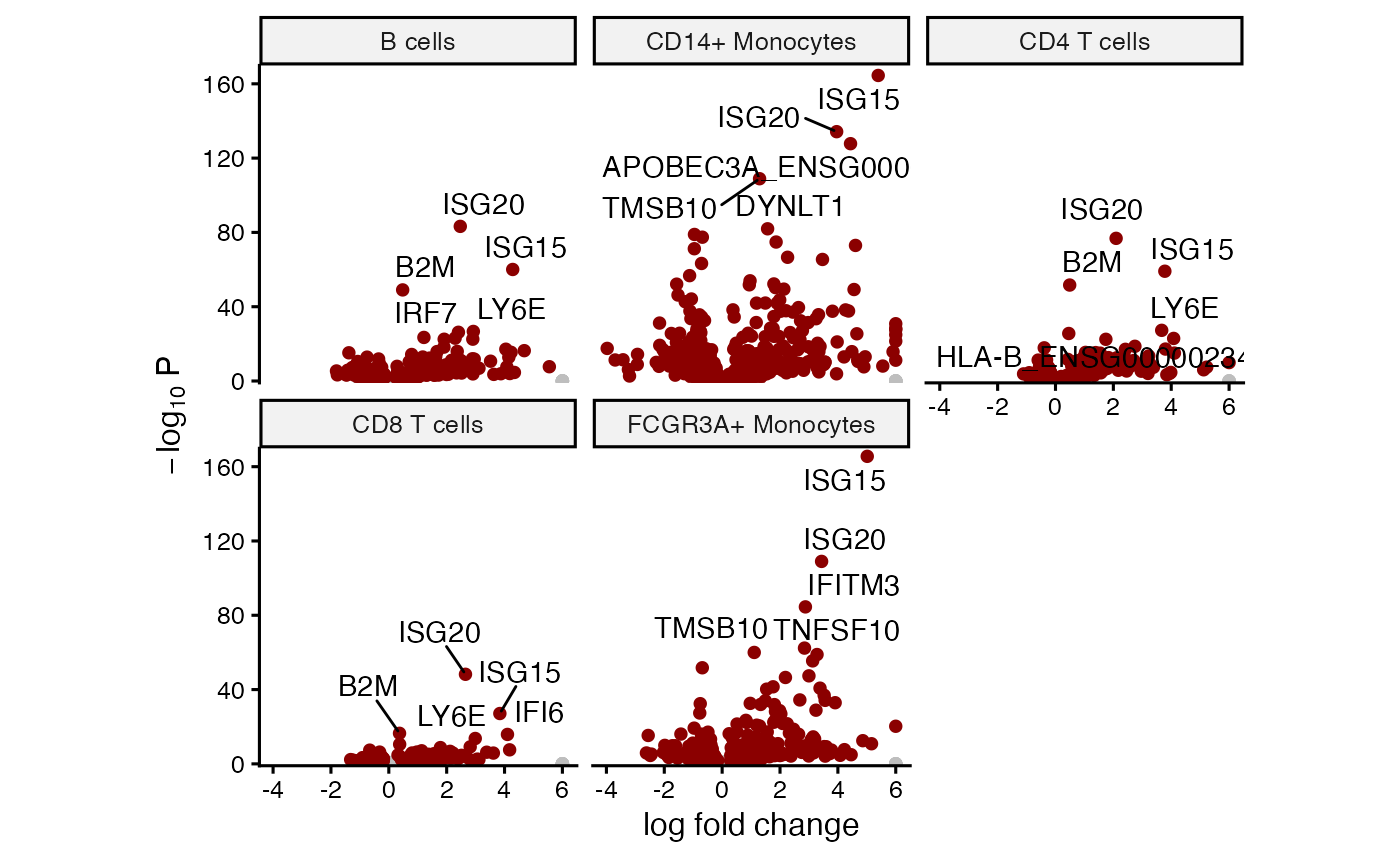
Volcano plot for each cell type
Usage
plotVolcano(
x,
coef,
method = c("classic", "Bayesian"),
nGenes = 5,
size = 12,
ymax = 300,
xmax = 6,
cutoff = 0.05,
ncol = 3,
cluster_ids = names(x),
...
)
# S4 method for class 'lucida'
plotVolcano(
x,
coef,
method = c("classic", "Bayesian"),
nGenes = 5,
size = 12,
ymax = 300,
xmax = 6,
cutoff = 0.05,
ncol = 3,
cluster_ids = names(x),
...
)Arguments
- x
result from
lucida()- coef
coefficient to test with
results()- method
"classic": multiple testing correction using Benjamini–Hochberg method, or "Bayesian": shrink coefficient estimates and report local FDR from
ashr::ash()using empirical Bayes approach.- nGenes
number of genes to highlight in each volcano plot
- size
text size
- ymax
maximum value to show on the y-axis
- xmax
maximum value to show on the x-axis
- cutoff
adj.P.Val cutoff to distinguish significant from non-significant genes
- ncol
number of columns in the plot
- cluster_ids
which cell types to plot
- ...
arguments passed to
facet_wrap(). Useful for specifyingscales = "free_y"
Examples
library(SingleCellExperiment)
# Load example data
data(example_sce, package="muscat")
sce <- example_sce
# Compute library size for each cell
sce$libSize <- colSums(counts(sce))
# Specify regression formula and cell annotation
form <- ~ group_id
fit <- lucida(sce, form, "cluster_id", verbose=FALSE)
#> B cells
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#> CD14+ Monocytes
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#> CD4 T cells
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#> CD8 T cells
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#> FCGR3A+ Monocytes
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
# Volcano plot
plotVolcano(fit, "group_idstim")