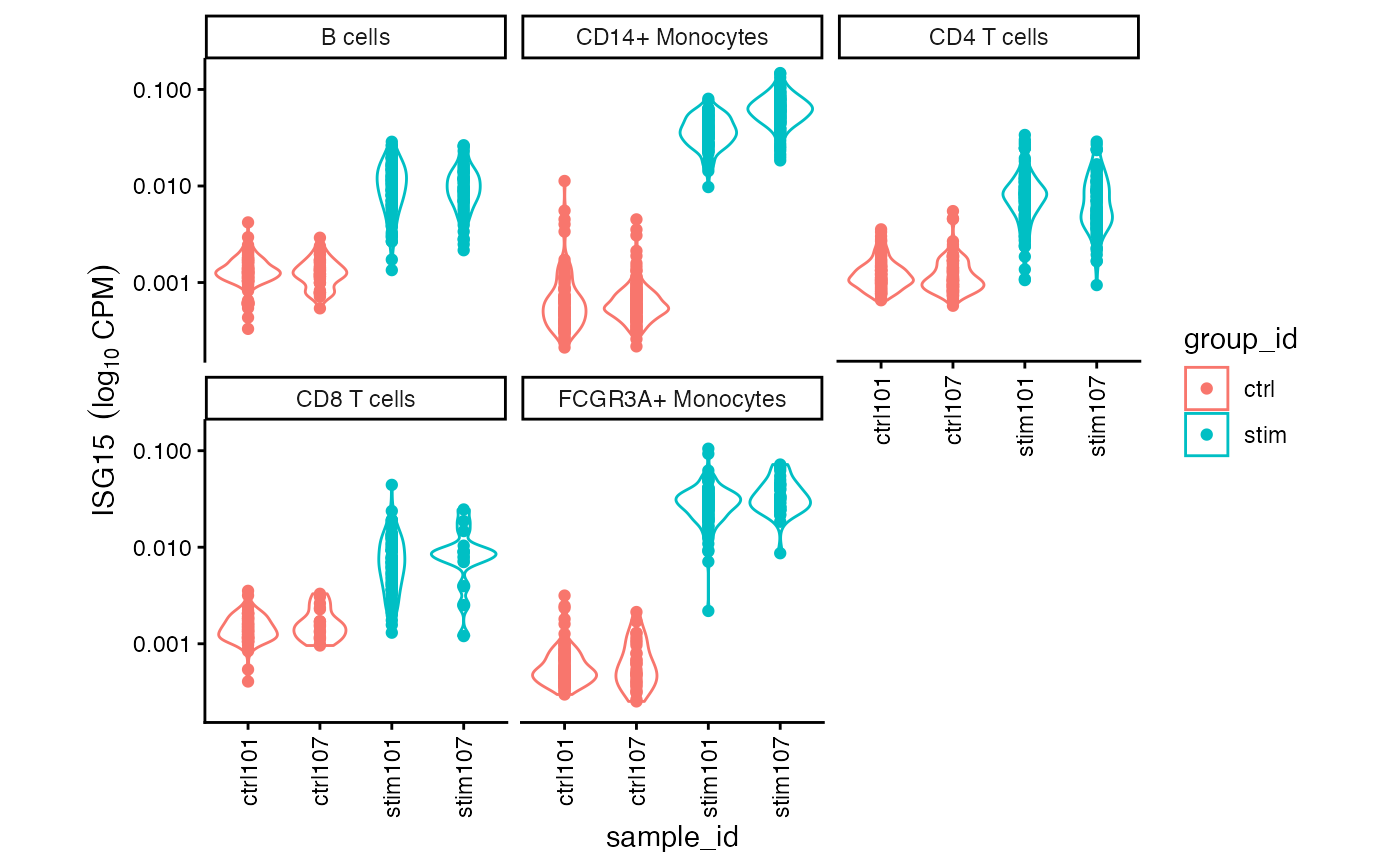
Plot expression stratified by cell type and trait
Usage
plotExpression(
sce,
gene,
sample_id,
cluster_id,
group_id,
cluster_ids = unique(sce[[cluster_id]])
)Examples
library(SingleCellExperiment)
# Load example data
data(example_sce, package="muscat")
sce <- example_sce
# Compute library size for each cell
sce$libSize <- colSums(counts(sce))
# Specify regression formula and cell annotation
form <- ~ group_id
fit <- lucida(sce, form, "cluster_id", verbose=FALSE)
#> B cells
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#> CD14+ Monocytes
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#> CD4 T cells
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#> CD8 T cells
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#> FCGR3A+ Monocytes
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
#>
| | 0%, ETA NA
|=======================================================| 100%, Elapsed 00:00
# Expression plot
plotExpression( sce, "ISG15", "sample_id", "cluster_id", "group_id")