Multi-ancestry eQTL meta-analysis of human brain identifies candidate causal variants for brain-related traits
Abstract
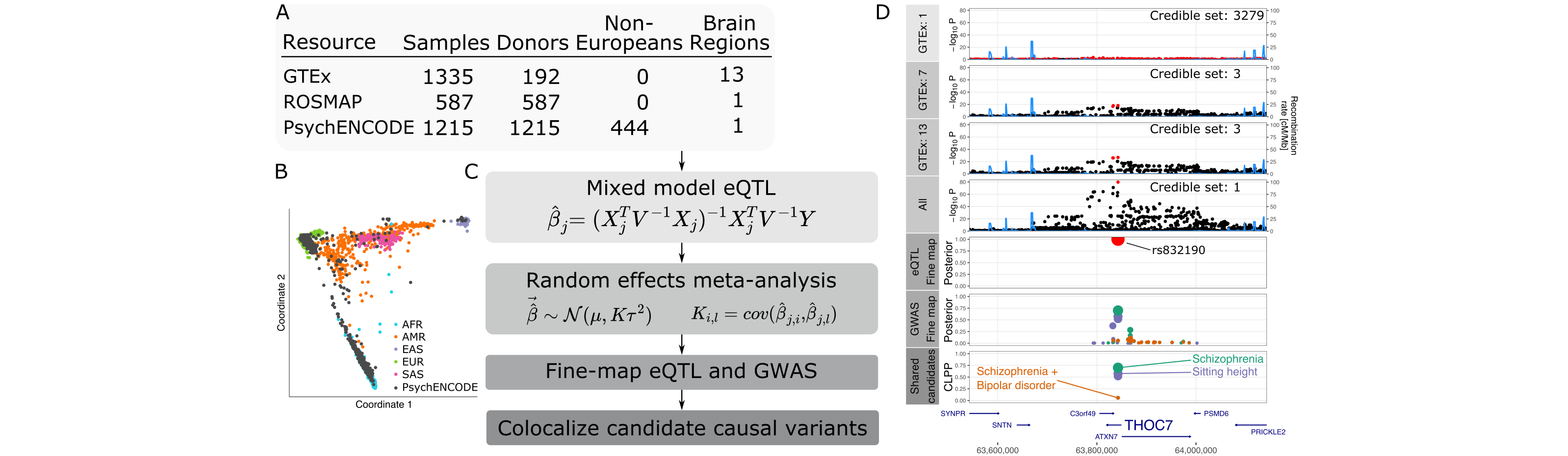
While large-scale, genome-wide association studies (GWAS) have identified hundreds of loci associated with brain-related traits, identification of the variants, genes and molecular mechanisms underlying these traits remains challenging. Integration of GWAS with expression quantitative trait loci (eQTLs) and identification of shared genetic architecture have been widely adopted to nominate genes and candidate causal variants. However, this approach is limited by sample size, statistical power and linkage disequilibrium. We developed the multivariate multiple QTL approach and performed a large-scale, multi-ancestry eQTL meta-analysis to increase power and fine-mapping resolution. Analysis of 3,983 RNA-sequenced samples from 2,119 donors, including 474 non-European individuals, yielded an effective sample size of 3,154. Joint statistical fine-mapping of eQTL and GWAS identified 329 variant–trait pairs for 24 brain-related traits driven by 204 unique candidate causal variants for 189 unique genes. This integrative analysis identifies candidate causal variants and elucidates potential regulatory mechanisms for genes underlying schizophrenia, bipolar disorder and Alzheimer’s disease.